 Select the IRS and PI3K group nodes by clicking on the group node outline.Once the nodes are grouped, we can add and style the interactiosn between them. The gene set now includes three group nodes. Repeat with the other complexes and groups, naming them PI3K and AKT respectively.Right-click and in the context menu, select Group → Group Selected Nodes.Select the three IRS nodes by clicking and dragging.Add Group Nodes to the Gene SetĬytoscape allows for grouping of nodes, which we can use to organize complexes and groups. In the gene set these have been expanded to include all members of isoforms and complex subunits. In this example, the original figure contained several complexes and isoform groups that were illustrated as single nodes. Use the symbol column to match nodes to the figure. Note that the label displayed on the node, name, is the official gene symbol, whereas the original published figures sometimes use various gene symbol aliases. We can start by moving the nodes into the approximate location from the figure. Next, we will arrange the gene set nodes and add interaction with the goal of reproducing the information from pathway figure in a computational model. Once the annotation is placed, enable annotation selection by clicking the Toggle Annotation Selection button in the Network View Tools under the network.Select the pathway figure you downloaded previously in the file browser.The initial placement can easily be changed later. In the Annotations tab of the Control Panel, select the Image icon at the top and then click anywhere in the network view to place the annotation.We can add the original published figure for this gene set as a Cytoscape annotation, to add context to the nodes. Set the Vertical spacing between nodes to 50 and Horizontal spacing between nodes to 80.Under Layout → Settings., select the Grid Layout. The gene set is already arranged with a grid layout, but we can make it a bit more compact.Set the default node fill color to light gray.In the Style interface, under Node Fill Color, create a continuous mapping for logFC, with the default ColorBrewer Red-Blue gradient.We can now add a simple visualization of the data on the set of nodes. Two new columns of data will be added to the Node Table. Select the ncbigene column as the Key column for Network, to match the Gene column of the data.Alternatively, drag and drop the data file directly onto the Node Table. Load the TCGA-Ovarian-MesenvsImmuno_data.csv file under File menu, select Import → Table from File. 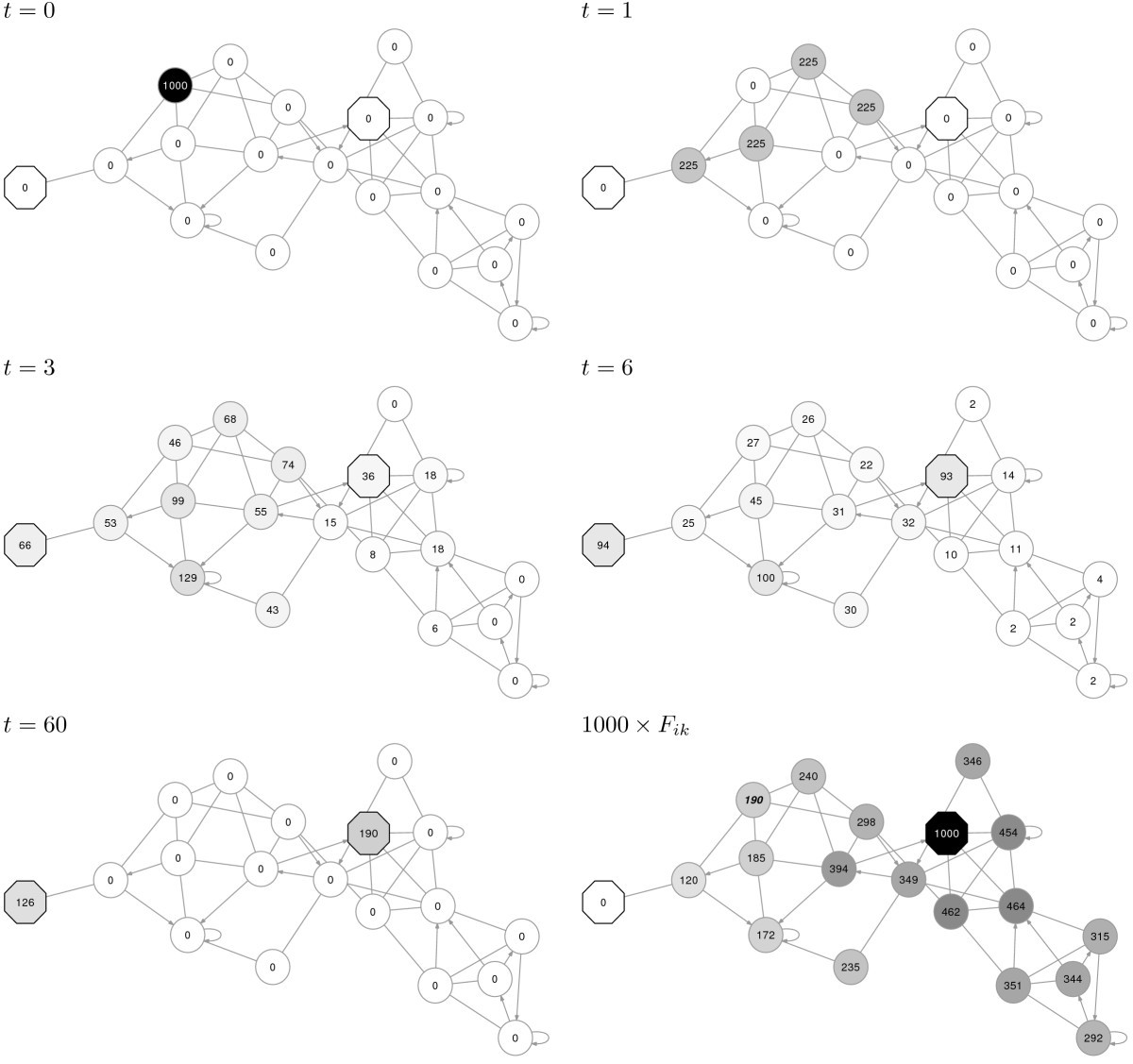
Download a local copy of TCGA-Ovarian-MesenvsImmuno_data.csv.The data is annotated with NCBI Gene identifiers, which is also what the nodes in the gene set are annotated with. Next, we will import experimental data from TCGA, comparing gene expression between two molecular subtypes of ovarian cancer (Mesenchymal and Immunoreactive). 
The gene set will appear as a grid of nodes in Cytoscape, similar to what it looks like at NDEx.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
